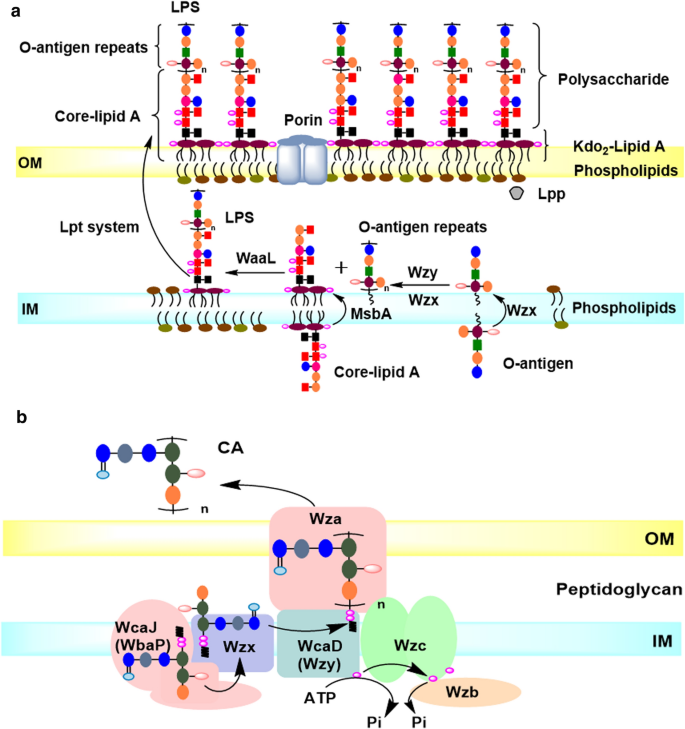
Insights into the structure of Escherichia coli outer membrane as the target for engineering microbial cell factories | Microbial Cell Factories | Full Text
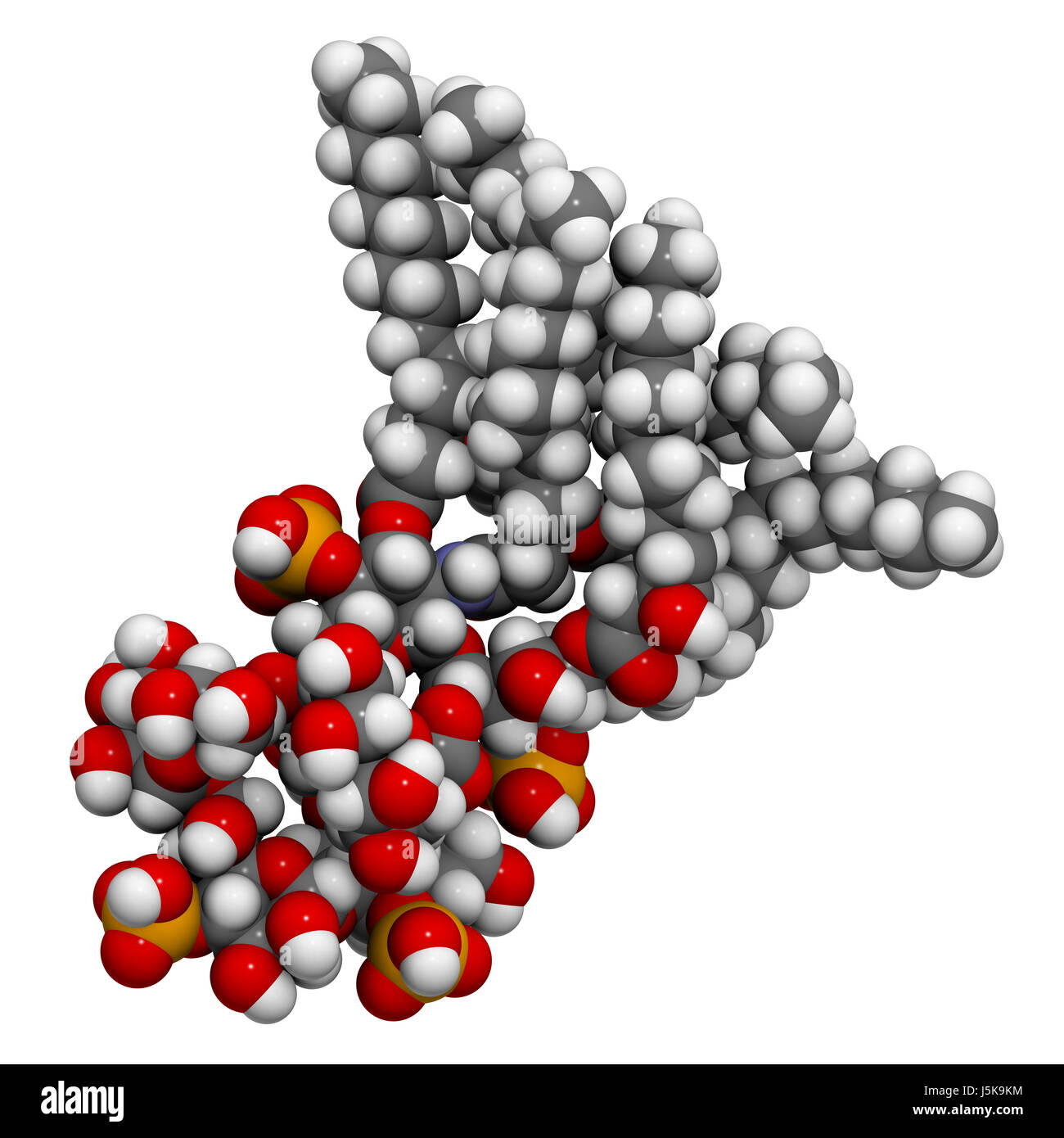
Lipopolysaccharide (LPS, lipid A and inner core fragment) endotoxin molecule from E. coli. 3D rendering based on protein data bank entry 3fxi Stock Photo - Alamy

Structural Characterization of a Model Gram-Negative Bacterial Surface Using Lipopolysaccharides from Rough Strains of Escherichia coli | Biomacromolecules
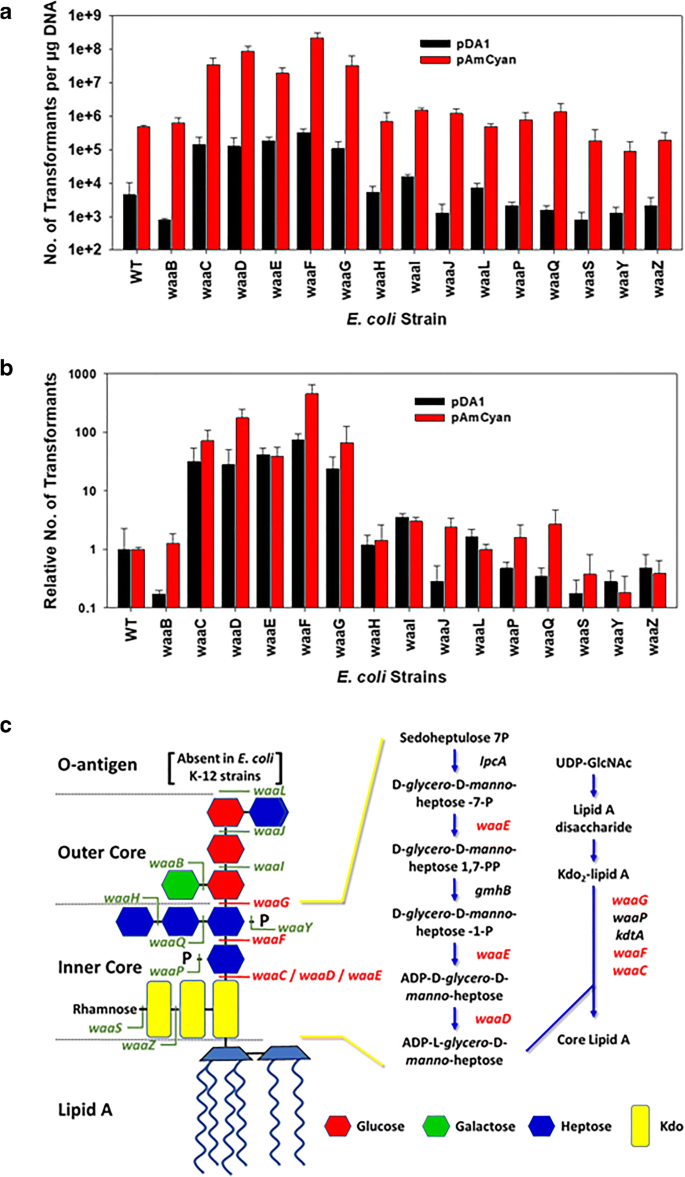
Loss of the lipopolysaccharide (LPS) inner core increases the electrocompetence of Escherichia coli | SpringerLink

General structure of E. coli LPS. The sugar moieties in the core region... | Download Scientific Diagram
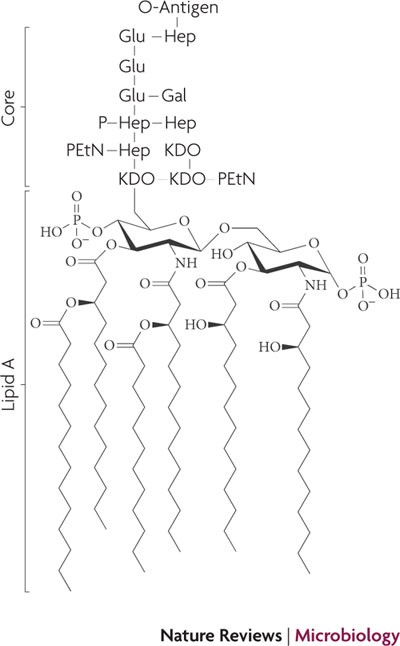
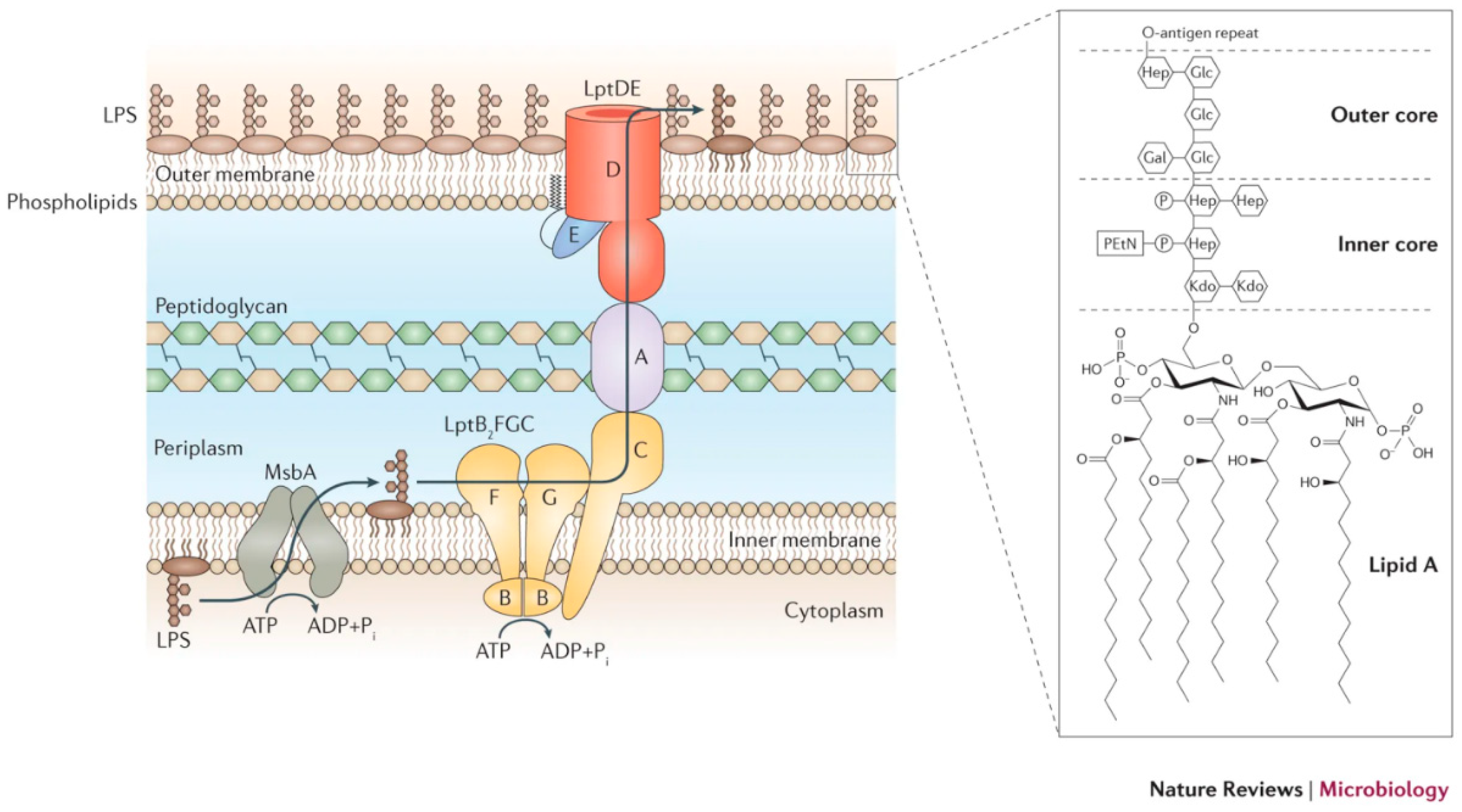
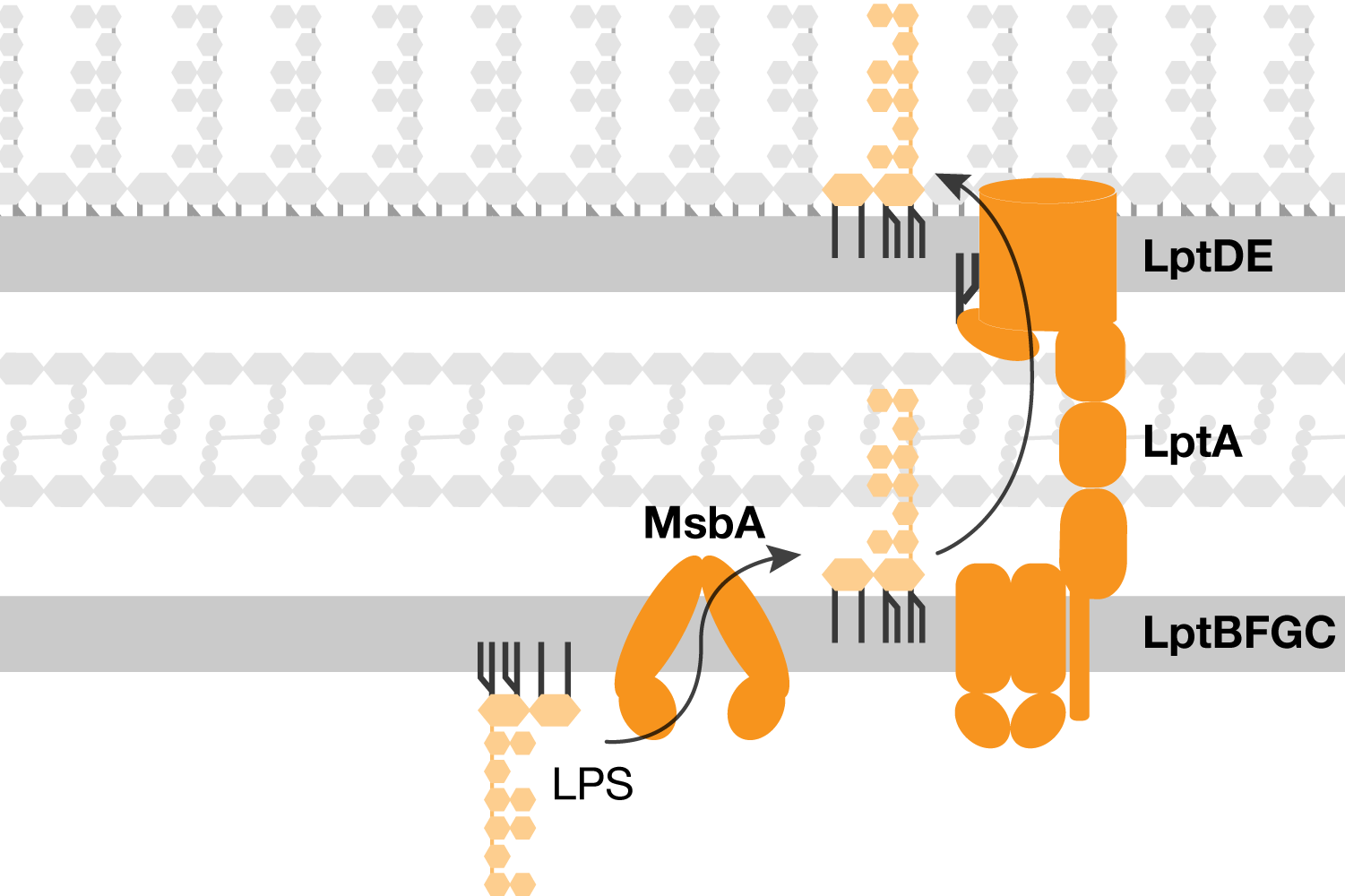
Transport of lipopolysaccharide across the cell envelope: the long road of discovery | Nature Reviews Microbiology

Detoxifying Escherichia coli for endotoxin-free production of recombinant proteins | Microbial Cell Factories | Full Text

Leveraging microfluidic dielectrophoresis to distinguish compositional variations of lipopolysaccharide in E. coli | bioRxiv

Enzymatic Synthesis of Lipopolysaccharide in Escherichia coli: PURIFICATION AND PROPERTIES OF HEPTOSYLTRANSFERASE I - ScienceDirect

Mutation and Suppressor Analysis of the Essential Lipopolysaccharide Transport Protein LptA Reveals Strategies To Overcome Severe Outer Membrane Permeability Defects in Escherichia coli | Journal of Bacteriology

E. coli-ΦX174 genotype to phenotype map reveals flexibility and diversity in LPS structures | bioRxiv
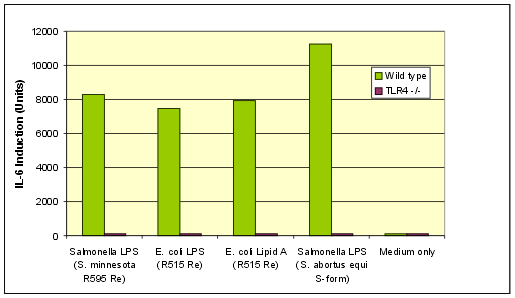
Lipid A from E. coli, Serotype R515 (Re) (TLRGRADE®) (Ready-to-Use) - ALX-581-200 - Enzo Life Sciences
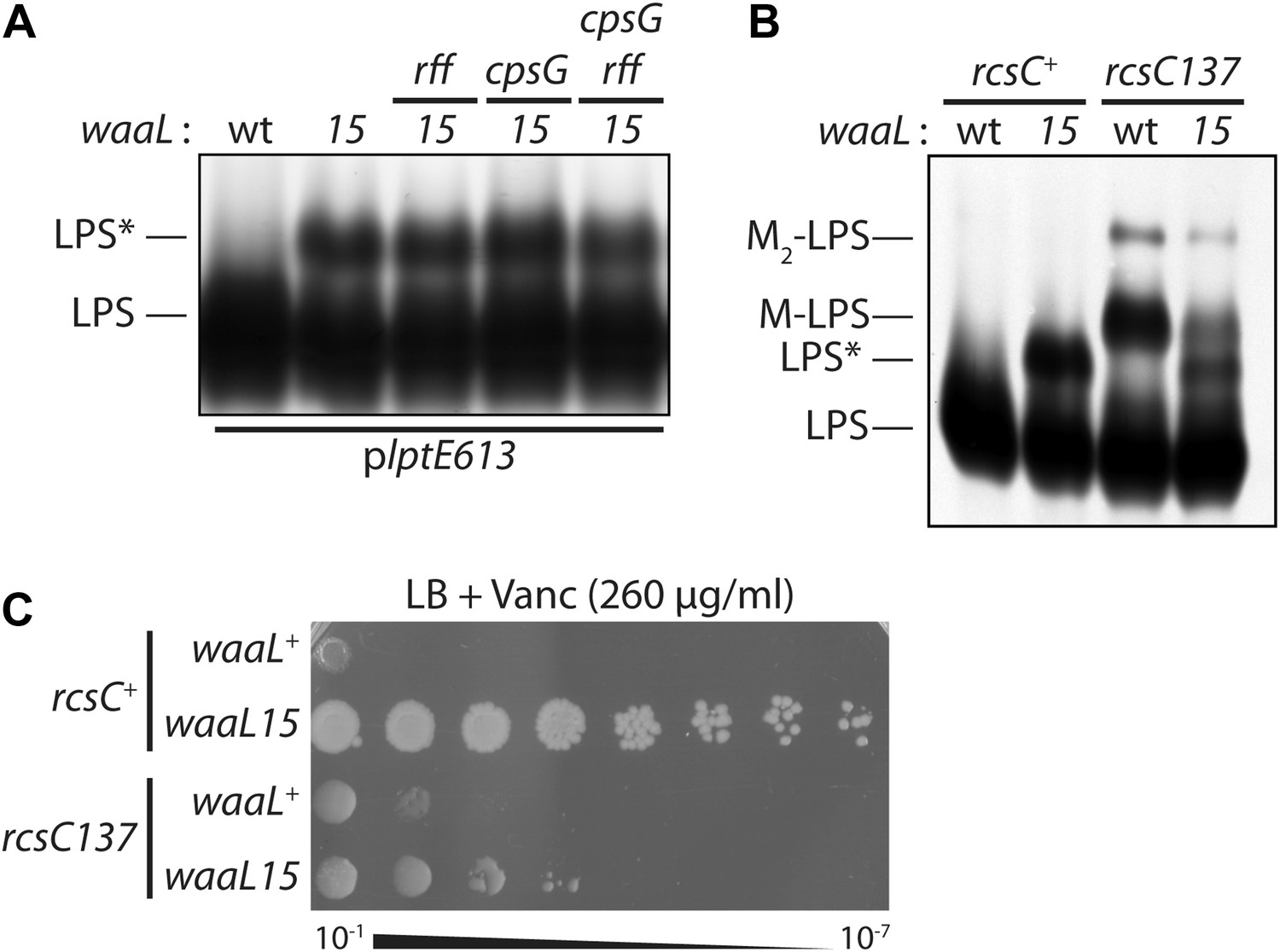
A mutant Escherichia coli that attaches peptidoglycan to lipopolysaccharide and displays cell wall on its surface | eLife
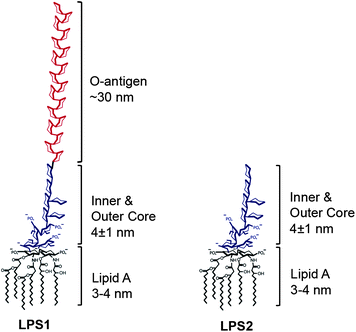
Understanding the molecular interactions of lipopolysaccharides during E. coli initial adhesion with a surface forces apparatus - Soft Matter (RSC Publishing) DOI:10.1039/C1SM05554B

DE10013539B4 - Escherichia coli lipopolysaccharides, process for their preparation and their uses - Google Patents






![Anti-E. coli LPS antibody [2D7/1] (ab35654) | Abcam Anti-E. coli LPS antibody [2D7/1] (ab35654) | Abcam](https://www.abcam.com/ps/products/35/ab35654/Images/ab35654-48358-anti-e-coli-lps-antibody-2d7-1-immunofluorescence.jpg)