PLOS Biology: ClipKIT: A multiple sequence alignment trimming software for accurate phylogenomic inference
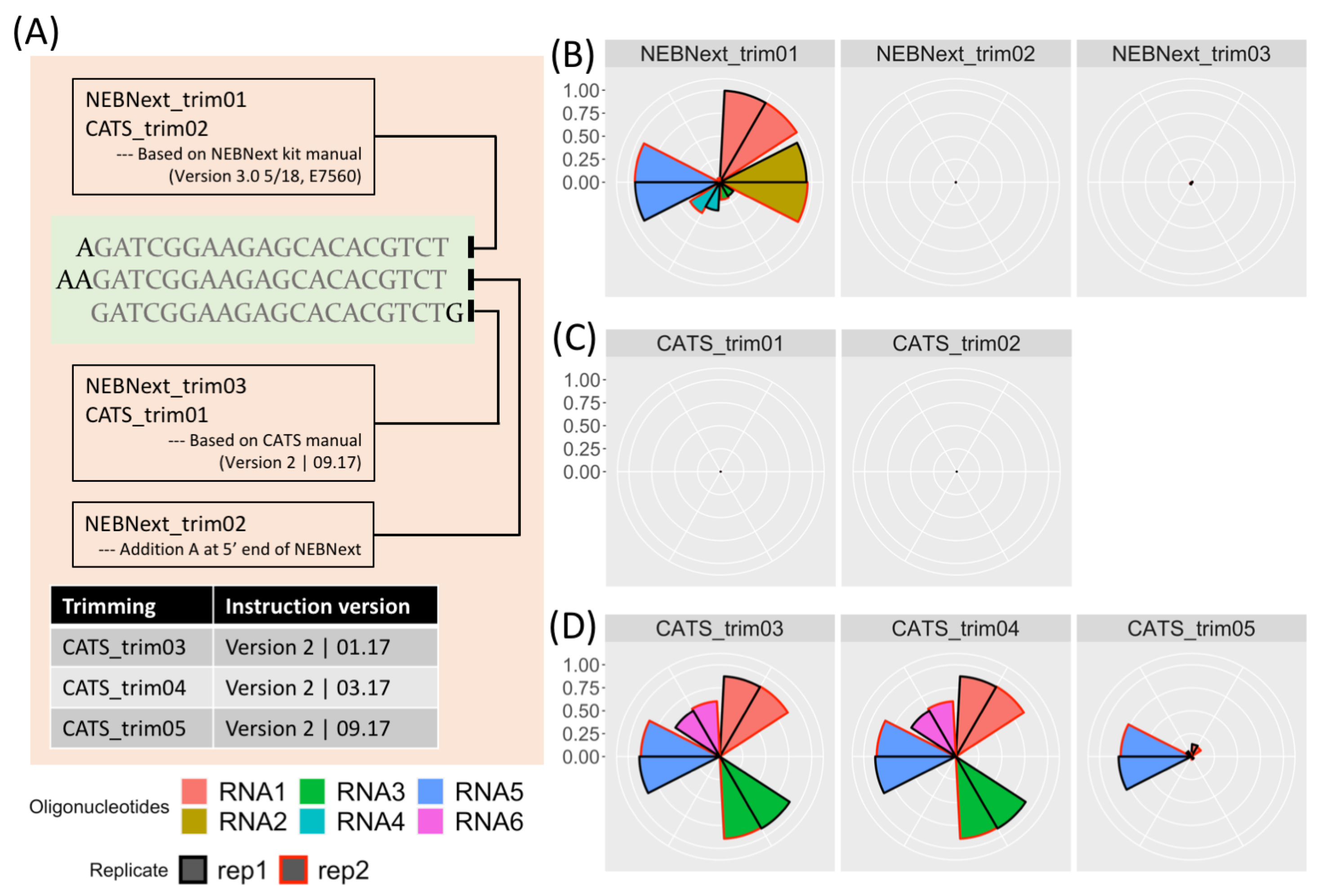
ncRNA | Free Full-Text | Accurate Adapter Information Is Crucial for Reproducibility and Reusability in Small RNA Seq Studies | HTML
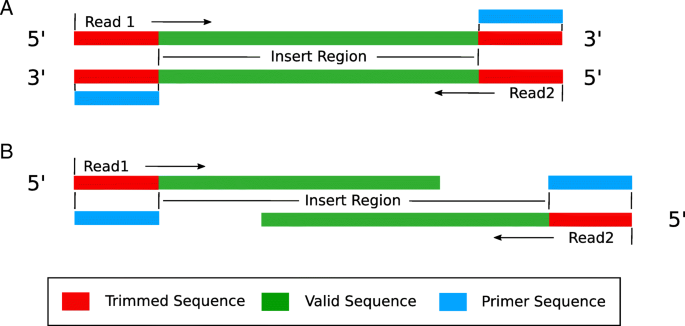
pTrimmer: An efficient tool to trim primers of multiplex deep sequencing data | BMC Bioinformatics | Full Text
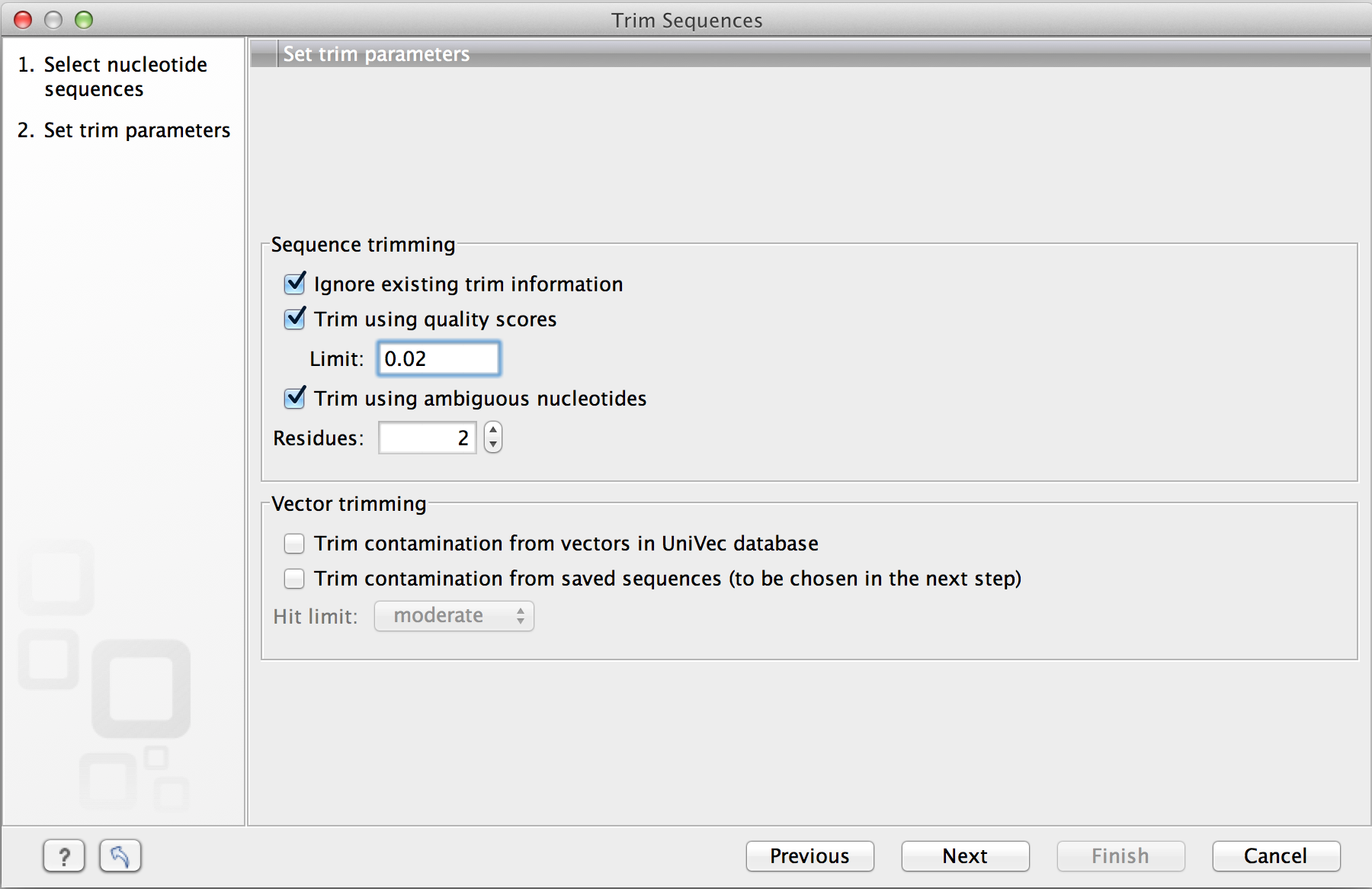
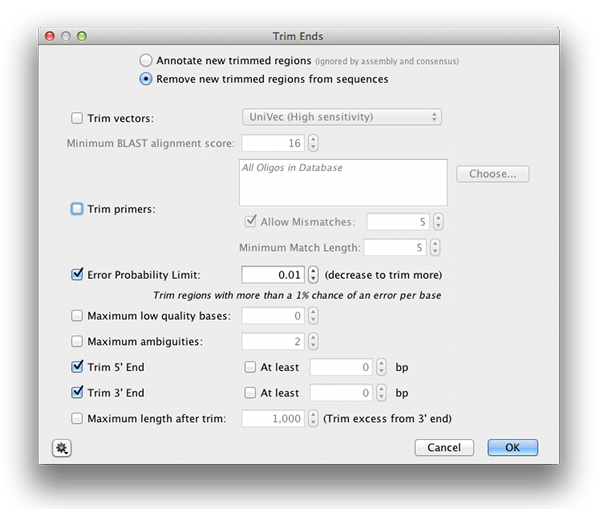
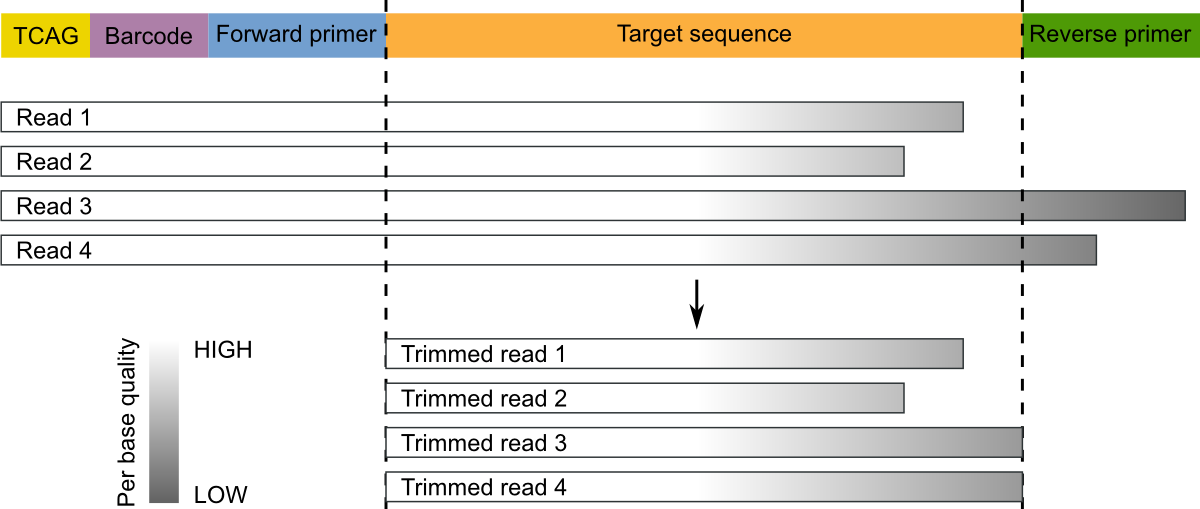
Automatic trimming Sequence reads can be trimmed automatically based on a number of different criteria. Automatic trimming is particularly useful in the following situations: If you have many sequence reads to be trimmed. If you wish to trim vector ...

Adapter trimming for single end sRNA-seq data. The reads processed by... | Download Scientific Diagram




![SEQUENCE [SEQ1-27] - $0.50 : eeagal Trimming! SEQUENCE [SEQ1-27] - $0.50 : eeagal Trimming!](https://www.eeagaltrimming.com/images/SEQ1-27.jpg)